Note
Run this example online :
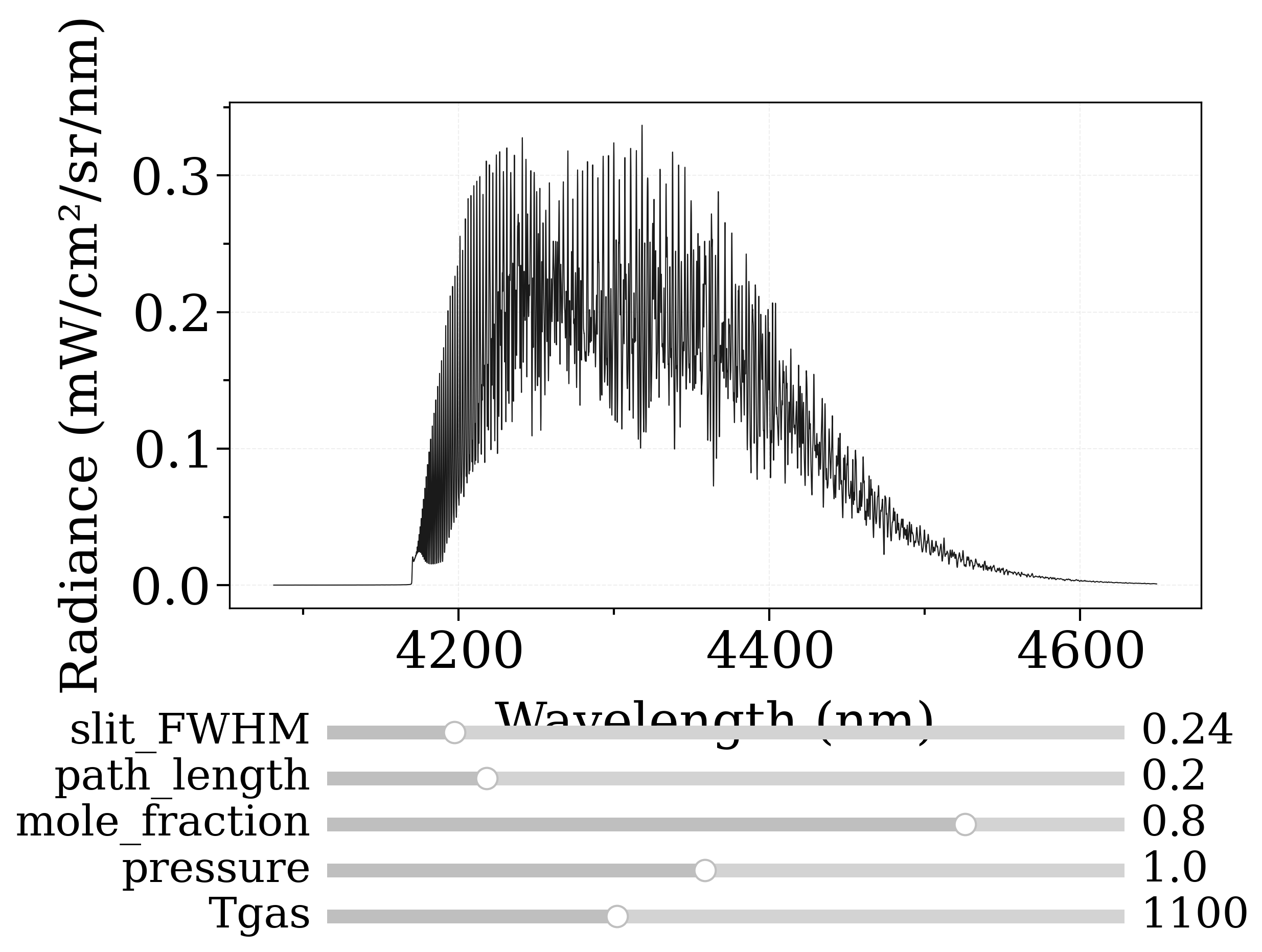
Real-time GPU Accelerated Spectra (Interactive)¶
Example using GPU sliders and GPU calculation with eq_spectrum_gpu()
This method requires CUDA compatible hardware to execute. For more information on how to setup your system to run GPU-accelerated methods using CUDA and Cython, check GPU Spectrum Calculation on RADIS
Note
in the example below, the GPU code runs on CPU, using the parameter emulate=True.
In your environment, to run the GPU code with the full power of the GPU, remove this line
or set emulate=False (default)
HITEMP keep only relevant input files: ['/home/docs/.radisdb/hitemp/CO2-02_02125-02250_HITEMP2010.hdf5', '/home/docs/.radisdb/hitemp/CO2-02_02250-02500_HITEMP2010.hdf5']
HITEMP keep only relevant input files: []
HITEMP keep only relevant input files: ['/home/docs/.radisdb/hitemp/CO2-02_02125-02250_HITEMP2010.hdf5', '/home/docs/.radisdb/hitemp/CO2-02_02250-02500_HITEMP2010.hdf5']
Number of lines loaded: 1128265
Finished calculating spectrum!
1.19s - Spectrum calculated
Spectrum Name: CO2-hitemp-radisdb-1100.0K-#1168
Spectral Quantities
----------------------------------------
abscoeff [cm-1] (150,001 points)
absorbance (150,001 points)
emissivity (150,001 points, 242 nans)
emissivity_noslit (150,001 points)
transmittance_noslit (150,001 points)
radiance_noslit [mW/cm2/sr/cm-1] (150,001 points)
transmittance (150,001 points, 242 nans)
radiance [mW/cm2/sr/cm-1] (150,001 points, 242 nans)
Physical Conditions
----------------------------------------
Tgas 1100.0 K
Trot 1100.0 K
Tvib 1100.0 K
isotope 1,2,3
mole_fraction 0.8
molecule CO2
overpopulation None
path_length 0.2 cm
pressure_mbar 1000.0 mbar
rot_distribution boltzmann
self_absorption True
state X
thermal_equilibrium True
vib_distribution boltzmann
wavenum_max 2450.0000 cm-1
wavenum_min 2150.0000 cm-1
Computation Parameters
----------------------------------------
NwG 3
NwL 6
Tref 296 K
add_at_used
broadening_method voigt
cutoff 0 cm-1/(#.cm-2)
dbformat hitemp-radisdb
dbpath /home/docs/.radisdb/hitemp/CO2-02_02125-02250_HITEMP2010.hdf5,/home/docs/.radisdb/hitemp/CO2-02_0225...
default_output_unit cm-1
emulate_gpu True
folding_thresh 1e-06
include_neighbouring_lines True
memory_mapping_engine auto
neighbour_lines 0 cm-1
norm_by area
optimization simple
parfuncfmt hapi
parsum_mode full summation
pressure 1.0
profiler {'spectrum_calculation': {'scaled_S0': 0.03147624399571214, 'calc_other_spectral_quan': 0.0132299820...
pseudo_continuum_threshold 0
radis_version 0.13.1
slit_FWHM 0.24
slit_dispersion None
slit_dispersion_threshold 0.01
slit_function 0.24
slit_shape triangular
slit_unit cm-1
sparse_ldm auto
spectral_points 150000.0
truncation 50 cm-1
waveunit cm-1
wstep 0.002 cm-1
zero_padding -1
Config parameters
----------------------------------------
DEFAULT_DOWNLOAD_PATH ~/.radisdb
GRIDPOINTS_PER_LINEWIDTH_ERROR_THRESHOLD 1
GRIDPOINTS_PER_LINEWIDTH_WARN_THRESHOLD 3
SPARSE_WAVERANGE auto
Information
----------------------------------------
calculation_time 1.192465706000803 s
chunksize None
db_use_cached True
dxG 0.13753507880165727
dxL 0.20180288881201608
export_lines False
export_populations None
export_rovib_fraction False
levelsfmt radis
lines_calculated 1,128,265
load_energies False
lvl_use_cached True
parfuncpath None
total_lines 1128265
warning_broadening_threshold 0.01
warning_linestrength_cutoff 0.01
wavenum_max_calc 2450.0000 cm-1
wavenum_min_calc 2150.0000 cm-1
----------------------------------------
from radis import SpectrumFactory
# from radis.test.utils import getTestFile
from radis.tools.plot_tools import ParamRange
## Add an experimental file :
# my_file = getTestFile("CO2_measured_spectrum_4-5um.spec") # for the example here
# s_exp = load_spec(my_file)
# s_exp.crop(4120, 4790).plot(Iunit="mW/cm2/sr/nm")
#
## This spectrum is significantly absorbed by atmospheric CO2
## so it will never match the synthetic spectrum.
## TODO: find different spectrum for this example.
sf = SpectrumFactory(
2150,
2450, # cm-1
molecule="CO2",
isotope="1,2,3",
wstep=0.002,
)
sf.fetch_databank("hitemp")
s = sf.eq_spectrum_gpu_interactive(
var="radiance",
Tgas=ParamRange(300.0, 2500.0, 1100.0), # K
pressure=ParamRange(0.1, 2, 1), # bar
mole_fraction=ParamRange(0, 1, 0.8),
path_length=ParamRange(0, 1, 0.2), # cm
slit_FWHM=ParamRange(0, 1.5, 0.24), # cm-1
emulate=True, # if True, runs CPU code on GPU. Set to False or remove to run on the GPU
plotkwargs={"wunit": "nm"}, # "nfig": "same",
)
print(s)
Total running time of the script: ( 0 minutes 2.881 seconds)



