Note
Run this example online :
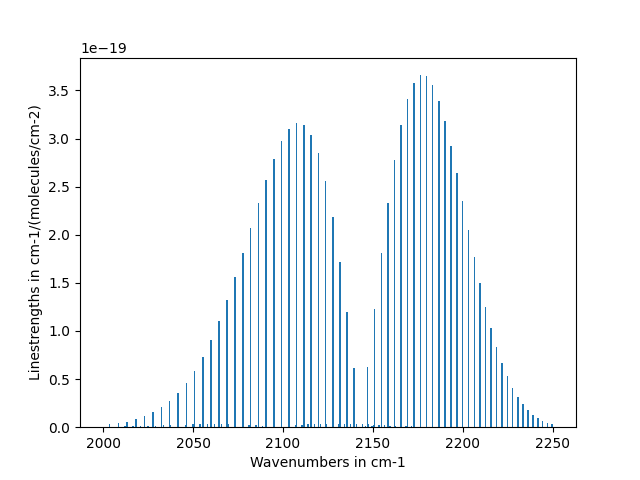
Scale Linestrengths of carbon-monoxide¶
This example scales the linestrengths of CO to Tgas=300 from Tref = 296 and then plots
the linestrengths against the wavenumbers. We are using fetch_hitemp()
function to retrieve the dataframe from the HITEMP-CO databank.
References¶
\[S(T) = S_0 \frac{Q_{ref}}{Q_{gas}} \operatorname{exp}\left(-E_l \left(\frac{1}{T_{gas}}-\frac{1}{T_{ref}}\right)\right) \frac{1-\operatorname{exp}\left(\frac{-\omega_0}{Tgas}\right)}{1-\operatorname{exp}\left(\frac{-\omega_0}{T_{ref}}\right)}\]
See Eq.(A11) in [Rothman-1998]
Similar functions are used directly at the hearth of RADIS’s SpectrumFactory in the
calc_linestrength_eq() and calc_linestrength_noneq() methods
Scaling equilibrium linestrength
from matplotlib import pyplot as plt
from numpy import exp
from radis.db.classes import get_molecule, get_molecule_identifier
from radis.io.hitemp import fetch_hitemp
from radis.levels.partfunc import PartFuncTIPS
from radis.phys.constants import hc_k
def get_Qgas(molecule, iso, T):
M = get_molecule_identifier(molecule)
Q = PartFuncTIPS(M, iso)
return Q.at(T=T)
def scale_linestrength_eq(df, Tref, Tgas):
print("Scaling equilibrium linestrength")
# %% Load partition function values
def _calc_Q(molecule, iso, T_ref, T_gas):
Qref = get_Qgas(molecule, iso, T_ref)
Qgas = get_Qgas(molecule, iso, T_gas)
return Qref, Qgas
id_set = df.id.unique()
id = list(id_set)[0]
molecule = get_molecule(id) # retrieve the molecule
iso_set = set(df.iso) # df1.iso.unique()
Qref_Qgas_ratio = {}
for iso in iso_set:
Qref, Qgas = _calc_Q(molecule, iso, Tref, Tgas)
Qref_Qgas_ratio[iso] = Qref / Qgas
# Scaling linestrength with the equations from Rotham's paper
line_strength = df.int * df["iso"].map(Qref_Qgas_ratio)
line_strength *= exp(-hc_k * df.El * (1 / Tgas - 1 / Tref))
line_strength *= (1 - exp(-hc_k * df.wav / Tgas)) / (1 - exp(-hc_k * df.wav / Tref))
# Add a fresh columns with the scaled linestrength
df["S"] = line_strength # [cm-1/(molecules/cm-2)]
# Just to make sure linestrength is indeed added
assert "S" in df
return df
if __name__ == "__main__":
Tref = 296
df = fetch_hitemp(
molecule="CO",
isotope="1, 2, 3",
load_wavenum_min=2000,
load_wavenum_max=2250,
)
Tgas = 450
df = scale_linestrength_eq(df, Tref, Tgas)
plt.bar(df["wav"], df["S"])
plt.xlabel("Wavenumbers in cm-1")
plt.ylabel("Linestrengths in cm-1/(molecules/cm-2)")
plt.show()
Total running time of the script: ( 0 minutes 5.720 seconds)



