radis.lbl.loader module¶
Summary¶
Module to host the databank loading / database initialisation parts of
SpectrumFactory. This is done through SpectrumFactory
inheritance of the DatabankLoader class defined here
Routine Listings¶
PUBLIC METHODS
radis.lbl.loader.DatabankLoader.load_databank()>>> load line databaseradis.lbl.loader.DatabankLoader.init_databank()>>> load loaderradis.lbl.loader.DatabankLoader.fetch_databank()>>> fetch from HITRAN onlineradis.lbl.loader.DatabankLoader.init_database()>>> to interact / generate a SpectrumDatabaseradis.lbl.loader.DatabankLoader.get_partition_function_interpolator()radis.lbl.loader.DatabankLoader.get_partition_function_calculator()
PRIVATE METHODS - DATABASE LOADING
radis.lbl.loader.DatabankLoader._load_databank()radis.lbl.loader.DatabankLoader._reload_databank()radis.lbl.loader.DatabankLoader._check_line_databank()radis.lbl.loader.DatabankLoader._retrieve_from_database()radis.lbl.loader.DatabankLoader._build_partition_function_interpolator()radis.lbl.loader.DatabankLoader._build_partition_function_calculator()
Most methods are written in inherited class with the following inheritance scheme:
DatabankLoader > BaseFactory >
BroadenFactory > BandFactory >
SpectrumFactory
.. inheritance-diagram:: radis.lbl.factory.SpectrumFactory
- parts
1
Notes
RADIS includes automatic rebuilding of Deprecated cache files + a global variable
to force regenerating them after a given version. See "OLDEST_COMPATIBLE_VERSION"
key in radis.config
——————————————————————————-
- class ConditionDict[source]¶
Bases:
dictA class to hold Spectrum calculation input conditions (
Input), computation parameters (Parameters), or miscalleneous parameters (MiscParams). Works like a dict except you can also access attribute with:v = a.key # equivalent to v = a[key]
Also can be copied, deepcopied, and parallelized in multiprocessing
Notes
for developers: Parameters and Input could also have simply derived from the (object) class, but it may have missed some convenients functions implemented for dict. For instance, how to be picked / unpickled.
See also
- class DatabankLoader[source]¶
Bases:
object
See also
- df0[source]¶
initial line database after loading. If for any reason, you want to manipulate the line database manually (for instance, keeping only lines emitting by a particular level), you need to access the
df0attribute ofSpectrumFactory.Warning
never overwrite the
df0attribute, else some metadata may be lost in the process. Only use inplace operations. If reducing the number of lines, add a df0.reset_index()- For instance::
- sf = SpectrumFactory(
wavenum_min= 2150.4, wavenum_max=2151.4, pressure=1, isotope=1)
sf.load_databank(‘HITRAN-CO-TEST’) sf.df0.drop(sf.df0[sf.df0.vu!=1].index, inplace=True) # keep lines emitted by v’=1 only sf.eq_spectrum(Tgas=3000, name=’vu=1’).plot()
df0contains the lines as they are loaded from the database.df1is generated during the spectrum calculation, after the line database reduction steps, population calculation, and scaling of intensity and broadening parameters with the calculated conditions.See also
df1- Type
pd.DataFrame
- df1[source]¶
line database, scaled with populations + linestrength cutoff Never edit manually. See all comments about
df0See also
df0- Type
pd.DataFrame
- fetch_databank(source='hitran', database='default', parfunc=None, parfuncfmt='hapi', levels=None, levelsfmt='radis', load_energies=False, include_neighbouring_lines=True, parse_local_global_quanta=True, drop_non_numeric=True, db_use_cached=True, lvl_use_cached=True, memory_mapping_engine='default', load_columns='equilibrium', parallel=True, extra_params=None)[source]¶
Fetch the latest files from [HITRAN-2020], [HITEMP-2010] (or newer), [ExoMol-2020] or [GEISA-2020] , and store them locally in memory-mapping formats for extremelly fast access.
- Parameters
source (
'hitran','hitemp','exomol','geisa') – which database to use.database (
'full','range', name of an ExoMol database, or'default') – if fetching from HITRAN,'full'download the full database and register it,'range'download only the lines in the range of the molecule.Note
'range'will be faster, but will require a new download each time you’ll change the range.'full'is slower and takes more memory, but will be downloaded only once.Default is
'full'.If fetching from HITEMP, only
'full'is available.if fetching from ‘’
exomol’’, use this parameter to choose which database to use. Keep'default'to use the recommended one. See all available databases withradis.io.exomol.get_exomol_database_list()By default, databases are download in
~/.radisdb. Can be changed inradis.config["DEFAULT_DOWNLOAD_PATH"]or in~/radis.jsonconfig file
- Other Parameters
parfuncfmt (
'cdsd','hapi','exomol', or any ofKNOWN_PARFUNCFORMAT) – format to read tabulated partition function file. If'hapi', then [HAPI] (HITRAN Python interface) is used to retrieve [TIPS-2020] tabulated partition functions. If'exomol'then partition functions are downloaded from ExoMol. Default'hapi'.parfunc (filename or None) – path to a tabulated partition function file to use.
levels (dict of str or None) –
path to energy levels (needed for non-eq calculations). Format:
{1:path_to_levels_iso_1, 3:path_to_levels_iso3}.
Default
Nonelevelsfmt (
'cdsd-pc','radis'(or any ofKNOWN_LVLFORMAT) orNone) – how to read the previous file. Known formats: (seeKNOWN_LVLFORMAT). Ifradis, energies are calculated using the diatomic constants in radis.db database if available for given molecule. Look up references there. IfNone, non equilibrium calculations are not possible. Default'radis'.load_energies (boolean) – if
False, dont load energy levels. This means that nonequilibrium spectra cannot be calculated, but it saves some memory. DefaultFalseinclude_neighbouring_lines (bool) – if
True, includes off-range, neighbouring lines that contribute because of lineshape broadening. Theneighbour_linesparameter is used to determine the limit. DefaultTrue.parse_local_global_quanta (bool, or
'auto') – ifTrue, parses the HITRAN/HITEMP ‘glob’ and ‘loc’ columns to extract quanta identifying the lines. Required for nonequilibrium calculations, or to useline_survey(), but takes up more space.drop_non_numeric (boolean) – if
True, non numeric columns are dropped. This improves performances, but make sure all the columns you need are converted to numeric formats before hand. DefaultTrue. Note that if a cache file is loaded it will be left untouched.db_use_cached (bool, or
'regen') – use cachedmemory_mapping_engine (
'pytables','vaex','feather') – which library to use to read HDF5 files (they are incompatible:'pytables'is row-major while'vaex'is column-major) or other memory-mapping formats If'default', use the value from ~/radis.json["MEMORY_MAPPING_ENGINE"]parallel (bool) – if
True, uses joblib.parallel to load database with multiple processes (works only for HITEMP files)load_columns (list,
'all','equilibrium','noneq') – columns names to load. If'equilibrium', only load the columns required for equilibrium calculations. If'noneq', also load the columns required for non-LTE calculations. Seedrop_all_but_these. If'all', load everything. Note that for performances, it is better to load only certain columsn rather than loading them all and dropping them withdrop_columns. Default'equilibrium'.Warning
if using
'equilibrium', not all parameters will be available for a Spectrumline_survey().
Notes
HITRAN is fetched with Astroquery [1]_ or [HAPI], and HITEMP with
fetch_hitemp()HITEMP files are generated in a ~/.radisdb database.See also
References
- get_abundance(molecule, isotope)[source]¶
Get isotopic abundance
- Parameters
molecule (str)
isotope (int, or list) – isotope number, sorted in terrestrial abundance
Examples
Use it from SpectrumFactory:
sf.get_abundance("H2O", 1) sf.get_abundance("CH4", [1,2,3])
See also
- get_conditions(ignore_misc=False, add_config=False)[source]¶
Get all parameters defined in the SpectrumFactory.
- Other Parameters
ignore_misc (boolean) – if
True, then all attributes considered as Factory ‘descriptive’ parameters, as defined inget_conditions()are ignored when comparing the database to current factory conditions. It should obviously only be attributes that have no impact on the Spectrum produced by the factory. DefaultFalse
- get_partition_function_calculator(molecule, isotope, elec_state)[source]¶
Retrieve Partition Function Calculator.
- Parameters
molecule (str)
isotope (int)
elec_state (str)
- get_partition_function_interpolator(molecule, isotope, elec_state)[source]¶
Retrieve Partition Function Interpolator.
- Parameters
molecule (str)
isotope (int)
elec_state (str)
- get_partition_function_molecule(molecule)[source]¶
Retrieve Partition Function for Molecule.
- Parameters
molecule (str)
- init_databank(*args, **kwargs)[source]¶
Method to init databank parameters but only load them when needed.
Databank is reloaded by
_check_line_databank()Same inputs Parameters asload_databank():- Parameters
name (a section name specified in your
~/radis.json) –.radishas to be created in your HOME (Unix) / User (Windows). If notNone, all other arguments are discarded. Note that all files in database will be loaded and it may takes some time. Better limit the database size if you already know what range you need. See Configuration file andDBFORMATfor expected~/radis.jsonformat- Other Parameters
path (str, list of str, None) – list of database files, or name of a predefined database in the Configuration file (
json) Accepts wildcards*to select multiple filesformat (
'hitran','cdsd-hitemp','cdsd-4000', or any ofKNOWN_DBFORMAT) – database type.'hitran'for HITRAN/HITEMP,'cdsd-hitemp'and'cdsd-4000'for the different CDSD versions. Default'hitran'parfuncfmt (
'hapi','cdsd', or any ofKNOWN_PARFUNCFORMAT) – format to read tabulated partition function file. Ifhapi, then HAPI (HITRAN Python interface) [1]_ is used to retrieve them (valid if your database is HITRAN data). HAPI is embedded into RADIS. Check the version. If partfuncfmt is None thenhapiis used. Defaulthapi.parfunc (filename or None) – path to tabulated partition function to use. If
parfuncfmtishapithenparfuncshould be the link to the hapi.py file. If not given, then the hapi.py embedded in RADIS is used (check version)levels (dict of str or None) – path to energy levels (needed for non-eq calculations). Format: {1:path_to_levels_iso_1, 3:path_to_levels_iso3}. Default
Nonelevelsfmt (‘cdsd-pc’, ‘radis’ (or any of
KNOWN_LVLFORMAT) orNone) – how to read the previous file. Known formats: (seeKNOWN_LVLFORMAT). Ifradis, energies are calculated using the diatomic constants in radis.db database if available for given molecule. Look up references there. IfNone, non equilibrium calculations are not possible. Default'radis'.db_use_cached (boolean, or
None) – ifTrue, a pandas-readable csv file is generated on first access, and later used. This saves on the datatype cast and conversion and improves performances a lot. But! … be sure to delete these files to regenerate them if you happen to change the database. If'regen', existing cached files are removed and regenerated. It is also used to load energy levels from.h5cache file if exist. IfNone, the value given on Factory creation is used. DefaultNoneload_energies (boolean) – if
False, dont load energy levels. This means that nonequilibrium spectra cannot be calculated, but it saves some memory. DefaultTrueinclude_neighbouring_lines (bool) –
True, includes off-range, neighbouring lines that contribute because of lineshape broadening. Theneighbour_linesparameter is used to determine the limit. DefaultTrue.drop_columns (list) – columns names to drop from Line DataFrame after loading the file. Not recommended to use, unless you explicitely want to drop information (for instance if dealing with too large databases). If
[], nothing is dropped. If'auto', parameters considered unnecessary are dropped. Seedrop_auto_columns_for_dbformatanddrop_auto_columns_for_levelsfmt. Default'auto'.load_columns (list,
'all','equilibrium','noneq') – columns names to load. If'equilibrium', only load the columns required for equilibrium calculations. If'noneq', also load the columns required for non-LTE calculations. Seedrop_all_but_these. If'all', load everything. Note that for performances, it is better to load only certain columsn rather than loading them all and dropping them withdrop_columns. Default'equilibrium'.Warning
if using
'equilibrium', not all parameters will be available for a Spectrumline_survey().*Other arguments are related to how to open the files*
Notes
Useful in conjonction with
init_database()when dealing with large line databanks when some of the spectra may have been precomputed in a spectrum database (SpecDatabase) Note that any previously loaded databank is discarded on the method callSee also
-,-,-
- init_database(path, autoretrieve=True, autoupdate=True, add_info=['Tvib', 'Trot'], add_date='%Y%m%d', compress=True, **kwargs)[source]¶
Init a
SpecDatabasefolder inpathto later store our spectra. Spectra can also be automatically retrieved from the database instead of being calculated.- Parameters
path (str) – path to database folder. If it doesnt exist, create it Accepts wildcards
*to select multiple filesautoretrieve (boolean, or
'force') – ifTrue, a database lookup is performed whenever a new spectrum is calculated. If the spectrum already exists then it is retrieved from the database instead of being calculated. Spectra are considered the same if all the stored conditions fit. If set to'force', an error is raised if the spectrum is not found in the database (use it for debugging). DefaultTrueautoupdate (boolean) – if
True, all spectra calculated by this Factory are automatically exported in database. DefaultTrue(but only if init_database is explicitely called by user)add_info (list, or
None/False) – append these parameters and their values if they are in conditions. Default['Tvib', 'Trot']add_date (str, or
None/False) – adds date in strftime format to the beginning of the filename. Default ‘%Y%m%d’compress (boolean, or 2) – if
True, Spectrum are read and written in binary format. This is faster, and takes less memory space. DefaultTrue. If2, additionaly remove all redundant quantities.
- Other Parameters
**kwargs (**dict) – arguments sent to
SpecDatabaseinitialization.- Returns
db – the database where spectra will be stored or retrieved
- Return type
- load_databank(name=None, path=None, format: ['hitran', 'hitemp', 'cdsd-hitemp', 'cdsd-4000', 'hitemp-radisdb', 'hdf5-radisdb', 'geisa'] = None, parfunc=None, parfuncfmt: ['cdsd', 'hapi'] = None, levels=None, levelsfmt: ['radis', 'cdsd-pc', 'cdsd-pcN', 'cdsd-hamil', None] = None, db_use_cached=True, lvl_use_cached=True, load_energies=False, include_neighbouring_lines=True, drop_columns='auto', load_columns='equilibrium')[source]¶
Loads databank from shortname in the Configuration file. (
json), or by manually setting all attributes.Databank includes: - lines - partition function & format (tabulated or calculated) - (optional) energy levels, format
- Parameters
name (a section name specified in your
~/radis.json) –.radishas to be created in your HOME (Unix) / User (Windows). If notNone, all other arguments are discarded. Note that all files in database will be loaded and it may takes some time. Better limit the database size if you already know what range you need. See Configuration file andDBFORMATfor expected~/radis.jsonformat- Other Parameters
path (str, list of str, None) – list of database files, or name of a predefined database in the Configuration file (
json) Accepts wildcards*to select multiple filesformat (
'hitran','cdsd-hitemp','cdsd-4000', or any ofKNOWN_DBFORMAT) – database type.'hitran'for HITRAN/HITEMP,'cdsd-hitemp'and'cdsd-4000'for the different CDSD versions. Default'hitran'parfuncfmt (
'hapi','cdsd', or any ofKNOWN_PARFUNCFORMAT) – format to read tabulated partition function file. Ifhapi, then HAPI (HITRAN Python interface) [1]_ is used to retrieve them (valid if your database is HITRAN data). HAPI is embedded into RADIS. Check the version. If partfuncfmt is None thenhapiis used. Defaulthapi.parfunc (filename or None) – path to tabulated partition function to use. If
parfuncfmtishapithenparfuncshould be the link to the hapi.py file. If not given, then the hapi.py embedded in RADIS is used (check version)levels (dict of str or None) – path to energy levels (needed for non-eq calculations). Format: {1:path_to_levels_iso_1, 3:path_to_levels_iso3}. Default
Nonelevelsfmt (‘cdsd-pc’, ‘radis’ (or any of
KNOWN_LVLFORMAT) orNone) – how to read the previous file. Known formats: (seeKNOWN_LVLFORMAT). Ifradis, energies are calculated using the diatomic constants in radis.db database if available for given molecule. Look up references there. IfNone, non equilibrium calculations are not possible. Default'radis'.db_use_cached (boolean, or
None) – ifTrue, a pandas-readable csv file is generated on first access, and later used. This saves on the datatype cast and conversion and improves performances a lot. But! … be sure to delete these files to regenerate them if you happen to change the database. If'regen', existing cached files are removed and regenerated. It is also used to load energy levels from.h5cache file if exist. IfNone, the value given on Factory creation is used. DefaultTrueload_energies (boolean) – if
False, dont load energy levels. This means that nonequilibrium spectra cannot be calculated, but it saves some memory. DefaultTrueinclude_neighbouring_lines (bool) –
True, includes off-range, neighbouring lines that contribute because of lineshape broadening. Theneighbour_linesparameter is used to determine the limit. DefaultTrue.drop_columns (list) – columns names to drop from Line DataFrame after loading the file. Not recommended to use, unless you explicitely want to drop information (for instance if dealing with too large databases). If
[], nothing is dropped. If'auto', parameters considered useless are dropped. Seedrop_auto_columns_for_dbformatanddrop_auto_columns_for_levelsfmt. If'all', parameters considered unecessary for equilibrium calculations are dropped, including all information about lines that could be otherwise available inSpectrum()method. Warning: nonequilibrium calculations are not possible in this mode. Default'auto'.load_columns (list,
'all','equilibrium','noneq') – columns names to load. If'equilibrium', only load the columns required for equilibrium calculations. If'noneq', also load the columns required for non-LTE calculations. Seedrop_all_but_these. If'all', load everything. Note that for performances, it is better to load only certain columsn rather than loading them all and dropping them withdrop_columns. Default'equilibrium'.Warning
if using
'equilibrium', not all parameters will be available for a Spectrumline_survey().
See also
Only load when needed:
init_databank()Download from HITRAN:
fetch_databank()
Configuration file with: - all line database formats:
DBFORMAT- all energy levels database formats:LVLFORMATReferences
- misc[source]¶
Miscelleneous parameters (
MiscParams) params that cannot change the output of calculations (ex: number of CPU, etc.)
- molparam[source]¶
contains information about molar mass; isotopic abundance.
See
MolParams- Type
MolParam
- params[source]¶
Parametersthey may change the output of calculations (ex: threshold, cutoff, broadening methods, etc.)- Type
Computational parameters
- save_memory[source]¶
if True, tries to save RAM memory (but may take a little for time, saving stuff to files instead of RAM for instance)
- Type
bool
- set_abundance(molecule, isotope, abundance)[source]¶
Set isotopic abundance
- Parameters
molecule (str)
isotope (int, or list) – isotope number, sorted in terrestrial abundance
abundance (float, or list)
Examples
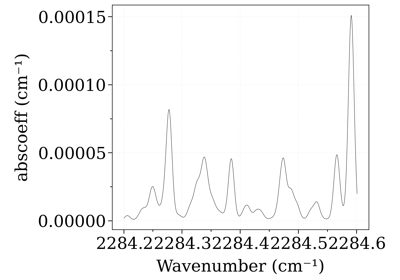
from radis import SpectrumFactory sf = SpectrumFactory( 2284.2, 2284.6, wstep=0.001, # cm-1 pressure=20 * 1e-3, # bar mole_fraction=400e-6, molecule="CO2", isotope="1,2", verbose=False ) sf.load_databank("HITEMP-CO2-TEST") print("Abundance of CO2[1,2]", sf.get_abundance("CO2", [1, 2])) sf.eq_spectrum(2000).plot("abscoeff") #%% Set the abundance of CO2(626) to 0.8; and the abundance of CO2(636) to 0.2 (arbitrary): sf.set_abundance("CO2", [1, 2], [0.8, 0.2]) print("New abundance of CO2[1,2]", sf.get_abundance("CO2", [1, 2])) sf.eq_spectrum(2000).plot("abscoeff", nfig="same")
See also
- warn(message, category='default', level=0)[source]¶
Trigger a warning, an error or just ignore based on the value defined in the
warningsdictionary. The warnings can thus be deactivated selectively by setting the SpectrumFactorywarningsattribute- Parameters
message (str) – what to print
category (str) – one of the keys of self.warnings. See
warningslevel (int) – warning level. Only print warnings when verbose level is higher than the warning levels. i.e., warnings of level 1 appear only if
verbose==True, warnings of level 2 appear only forverbose>=2, etc.. Warnings of level 0 appear only the time. Default0
Examples
- ::
- if not ((df.Erotu > tol).all() and (df.Erotl > tol).all()):
- self.warn(
“There are negative rotational energies in the database”, “NegativeEnergiesWarning”,
)
Notes
All warnings in the SpectrumFactory should call to this method rather than the default warnings.warn() method, because it allows easier runtime modification of how to deal with warnings
See also
- class Input[source]¶
Bases:
ConditionDictHolds Spectrum calculation input conditions, under the attribute
inputofSpectrumFactory. Works like a dict except you can also access attribute with:v = sf.input.key # equivalent to v = sf.input[key]
- KNOWN_DBFORMAT = ['hitran', 'hitemp', 'cdsd-hitemp', 'cdsd-4000', 'hitemp-radisdb', 'hdf5-radisdb', 'geisa'][source]¶
Known formats for Line Databases:
'hitran': [HITRAN-2020] original .par format.'hitemp': [HITEMP-2010] original format (same format as ‘hitran’).'cdsd-hitemp': CDSD-HITEMP original format (CO2 only, same lines as HITEMP-2010).'cdsd-4000': [CDSD-4000] original format (CO2 only).'hitemp-radisdb': HITEMP under RADISDB format (pytables-HDF5 with RADIS column names).'hdf5-radisdb': arbitrary HDF5 file with RADIS column names.'geisa': [GEISA-2020] original .par format.
To install all databases manually see the Configuration file and the list of databases .
See also
- Type
list
- KNOWN_LVLFORMAT = ['radis', 'cdsd-pc', 'cdsd-pcN', 'cdsd-hamil', None][source]¶
Known formats for Energy Level Databases (used in non-equilibrium calculations):
'radis': energies calculated with Dunham expansions by
'cdsd-pc': energies read from precomputed CDSD energies for CO2, withviblvl=(p,c)convention. SeePartFuncCO2_CDSDcalc
'cdsd-pcN': energies read from precomputed CDSD energies for CO2, withviblvl=(p,c,N)convention. SeePartFuncCO2_CDSDcalc
'cdsd-hamil': energies read from precomputed CDSD energies for CO2, withviblvl=(p,c,J,N)convention, i.e., a each rovibrational level can have a unique vibrational energy (this is needed when taking account Coupling terms) SeePartFuncCO2_CDSDcalc
None: means you can only do Equilibrium calculations.
See also
- Type
list
- KNOWN_PARFUNCFORMAT = ['cdsd', 'hapi'][source]¶
Known formats for partition function (tabulated files to read), or ‘hapi’ to fetch Partition Functions using HITRAN Python interface instead of reading a tabulated file.
See also
- Type
list
- class MiscParams[source]¶
Bases:
ConditionDictA class to hold Spectrum calculation descriptive parameters, under the attribute
paramsofSpectrumFactory. UnlikeParameters, these parameters cannot influence the Spectrum output and will not be used when comparing Spectrum with existing, precomputed spectra inSpecDatabaseWorks like a dict except you can also access attribute with:v = a.key
See also
input,params
- class Parameters[source]¶
Bases:
ConditionDictHolds Spectrum calculation computation parameters, under the attribute
paramsofSpectrumFactory. Works like a dict except you can also access attribute with:v = sf.params.key # equivalent to v = sf.params[key]
Also can be copied, deepcopied, and parallelized in multiprocessing
See also
input,misc- dbformat[source]¶
format of Line Database. See
KNOWN_DBFORMAT- Type
str
- include_neighbouring_lines[source]¶
if
True, includes the contribution of off-range, neighbouring lines because of lineshape broadening. DefaultTrue.- Type
bool
- levelsfmt[source]¶
format of Energy Database. See
KNOWN_LVLFORMAT- Type
str
- neighbour_lines[source]¶
extra range (cm-1) on each side of the spectrum to account for neighbouring lines. Overwritten by SpectrumFactory
- Type
float
- parfuncfmt[source]¶
format of tabulated Partition Functions. See #: str: format of Energy Database. See
KNOWN_PARFUNCFORMAT- Type
str
- parsum_mode[source]¶
“full summation” or “tabulation” . calculation mode of parittion function. See
RovibParFuncCalculator- Type
int
- pseudo_continuum_threshold[source]¶
threshold to assign lines in pseudo continuum. Overwritten in SpectrumFactory
- Type
float
- sparse_ldm[source]¶
“auto”, True, False . Sparse LDM calculation. See
radis.lbl.broadening.BroadenFactory._apply_lineshape_LDM()- Type
str
- truncation[source]¶
cutoff for half-width lineshape calculation (cm-1). Overwritten by SpectrumFactory
- Type
float
- wavenum_max_calc[source]¶
maximum calculated wavenumber (cm-1) initialized by SpectrumFactory
- Type
float
- df_metadata = ['molecule', 'iso', 'id', 'Ia', 'molar_mass', 'Qref', 'Qvib', 'Q', 'id', 'iso', 'id', 'id', 'id', 'id', 'id', 'iso', 'id', 'iso', 'id', 'id', 'id', 'iso', 'id', 'iso', 'id', 'id', 'id', 'iso', 'id', 'iso', 'id', 'iso', 'id', 'id', 'id', 'iso'][source]¶
metadata of line DataFrames
df0,df1. @dev: when having only 1 molecule, 1 isotope, these parameters are constant for all rovibrational lines. Thus, it’s faster and much more memory efficient to transport them as attributes of the DataFrame rather than columns. The syntax is the same, thus the operations do not change, i.e:k_b / df.molar_mass
will work whether molar_mass is a float or a column.
Warning
However, in the current Pandas implementation of
DataFrame, attributes are lost whenever the DataFrame is recreated, transposed, pickled.Thus, we use
transfer_metadata()to keep the attributes after an operation, andexpand_metadata()to make them columns before a Serializing operation (ex: multiprocessing) @dev: all of that is a high-end optimization. Users should not deal with internal DataFrames.References
https://stackoverflow.com/q/13250499/5622825
- Type
list
- drop_all_but_these = ['id', 'iso', 'wav', 'int', 'A', 'airbrd', 'selbrd', 'Tdpair', 'Tdpsel', 'Pshft', 'El', 'gp'][source]¶
drop all columns but these if using
drop_columns='all'in load_databank Note: nonequilibrium calculations wont be possible anymore and it wont be possible to identify lines withline_survey()See also
-,-,-- Type
list
- drop_auto_columns_for_dbformat = {'cdsd-4000': ['wang2'], 'cdsd-hitemp': ['wang2', 'lsrc'], 'geisa': [], 'hdf5-radisdb': [], 'hitemp': ['ierr', 'iref', 'lmix', 'gpp'], 'hitemp-radisdb': [], 'hitran': ['ierr', 'iref', 'lmix', 'gpp']}[source]¶
drop these columns if using
drop_columns='auto'in load_databank Based on the value ofdbformat=, some of these columns won’t be used.See also
-,-,-,-- Type
dict
- drop_auto_columns_for_levelsfmt = {'radis': [], 'cdsd-pc': ['v1u', 'v2u', 'l2u', 'v3u', 'ru', 'v1l', 'v2l', 'l2l', 'v3l', 'rl'], 'cdsd-pcN': ['v1u', 'v2u', 'l2u', 'v3u', 'ru', 'v1l', 'v2l', 'l2l', 'v3l', 'rl'], 'cdsd-hamil': ['v1u', 'v2u', 'l2u', 'v3u', 'ru', 'v1l', 'v2l', 'l2l', 'v3l', 'rl'], None: []}[source]¶
drop these columns if using
drop_columns='auto'in load_databank Based on the value oflvlformat=, some of these columns won’t be used.See also
-,-,-,-- Type
dict